
Assessing Escherichia coli metabolism models and simulation approaches in phenotype predictions: Validation against experimental data - Costa - 2018 - Biotechnology Progress - Wiley Online Library

Metabolite Labeling prediction. The combination of 13C labeling data... | Download Scientific Diagram
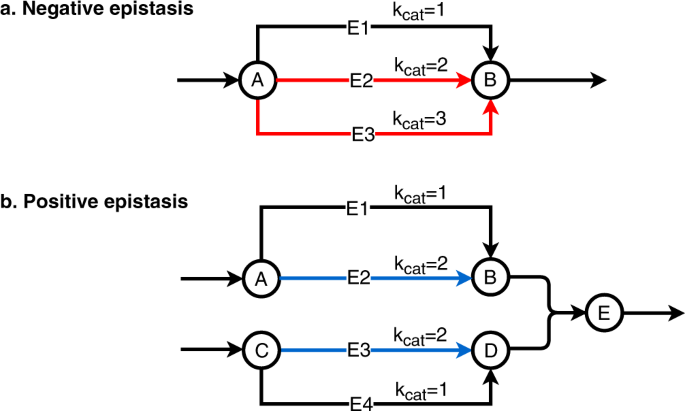
Flux balance analysis with or without molecular crowding fails to predict two thirds of experimentally observed epistasis in yeast | Scientific Reports
PLOS Computational Biology: Stoichiometric Representation of Gene–Protein–Reaction Associations Leverages Constraint-Based Analysis from Reaction to Gene-Level Phenotype Prediction
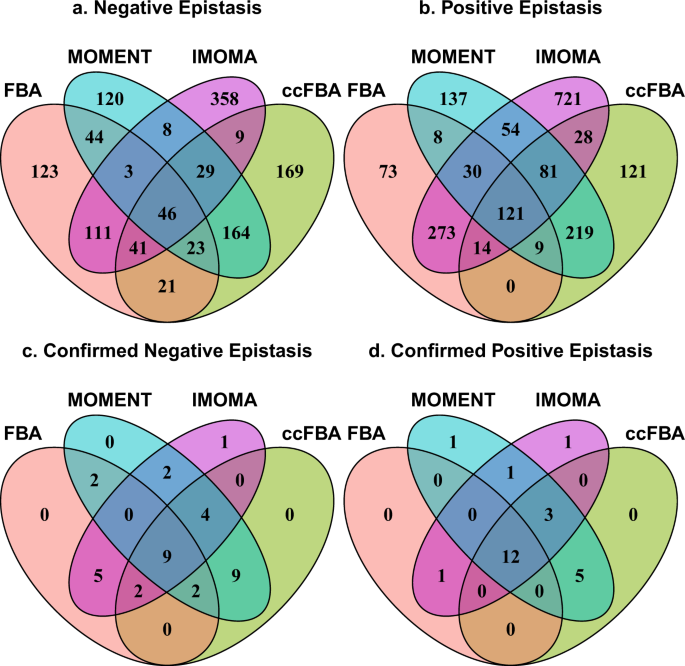
Flux balance analysis with or without molecular crowding fails to predict two thirds of experimentally observed epistasis in yeast | Scientific Reports

Grid-based computational methods for the design of constraint-based parsimonious chemical reaction networks to simulate metabolite production: GridProd | SpringerLink
Emergent sub-population behavior uncovered with a community dynamic metabolic model of Escherichia coli diauxic growth

Emergent Subpopulation Behavior Uncovered with a Community Dynamic Metabolic Model of Escherichia coli Diauxic Growth | mSystems
Emergent sub-population behavior uncovered with a community dynamic metabolic model of Escherichia coli diauxic growth Supplemen

Geometric illustration of the eMOMA method. Red-colored area represents... | Download Scientific Diagram
PLOS Computational Biology: In silico co-factor balance estimation using constraint-based modelling informs metabolic engineering in Escherichia coli
PLOS Computational Biology: Stoichiometric Representation of Gene–Protein–Reaction Associations Leverages Constraint-Based Analysis from Reaction to Gene-Level Phenotype Prediction

Improving the flux distributions simulated with genome-scale metabolic models of Saccharomyces cerevisiae - ScienceDirect
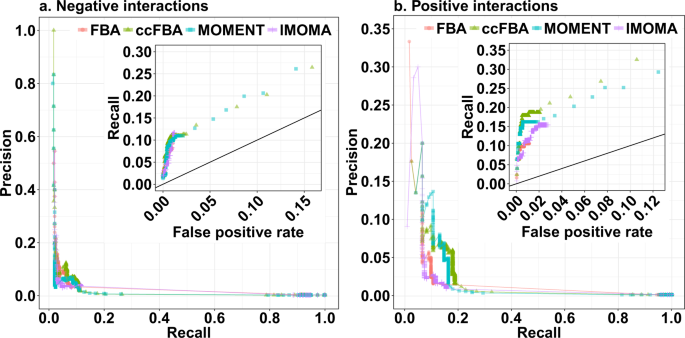
Flux balance analysis with or without molecular crowding fails to predict two thirds of experimentally observed epistasis in yeast | Scientific Reports

Improving the flux distributions simulated with genome-scale metabolic models of Saccharomyces cerevisiae - ScienceDirect

pFBA simulation of a single solution. Difference in simulation results... | Download Scientific Diagram

Assessing Escherichia coli metabolism models and simulation approaches in phenotype predictions: Validation against experimental data - Costa - 2018 - Biotechnology Progress - Wiley Online Library

Improving the flux distributions simulated with genome-scale metabolic models of Saccharomyces cerevisiae - ScienceDirect



![Full text] Designing metabolic engineering strategies with genome-scale metabolic | AGG Full text] Designing metabolic engineering strategies with genome-scale metabolic | AGG](https://www.dovepress.com/cr_data/article_fulltext/s58000/58494/img/Table1.jpg)